QuantumScale Single Cell RNA
Defy single cell gravity:
single cell at any scale
QuantumScale Single Cell RNA is powering the next generation of single cell with one flexible platform to meet any scale and research vision.
Benefits of QuantumScale Single Cell RNA
Ultra-efficient Workflow
Reduce hands-on time by over 75% without any specialized instrumentation
Flexible Project Sizes
Easily scale with a range of kit sizes for proof-of-concept studies to massive projects, from 168,000 to 4 million cells
Any Sample, Any Time
Compatible across species and cells and nuclei; lock in biology with fixation until you’re ready to run
Simplified Multiplexing
Easily combine 10 to >9,000 samples or conditions in a single run and eliminate batch effects with ScalePlex
Your Budget Goes Further
The most cost-effective solution on the market per cell, sample and experiment
Performance Maximized
High sensitivity and achieve cell recovery of 50%-60% or higher, while keeping multiplets low (≤4%)
View our AGBT 2025 workshop
Watch as our CEO Giovanna Prout presents Defy Single Cell Gravity: Escape platform constraints in single cell with QuantumScale.
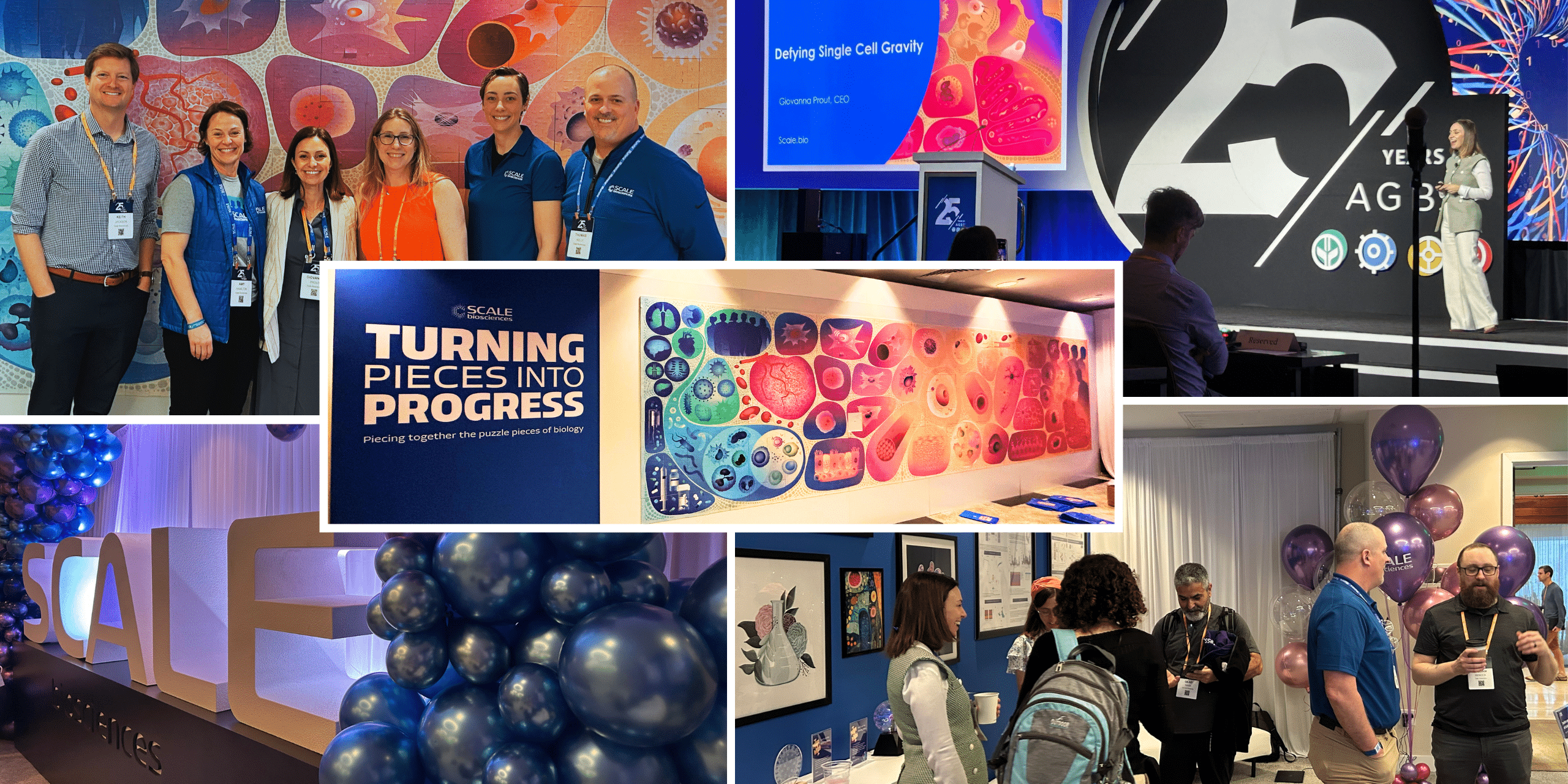


One solution for every scale
Your boldest projects, now possible. The most samples, now simpler than ever.
Key advantages
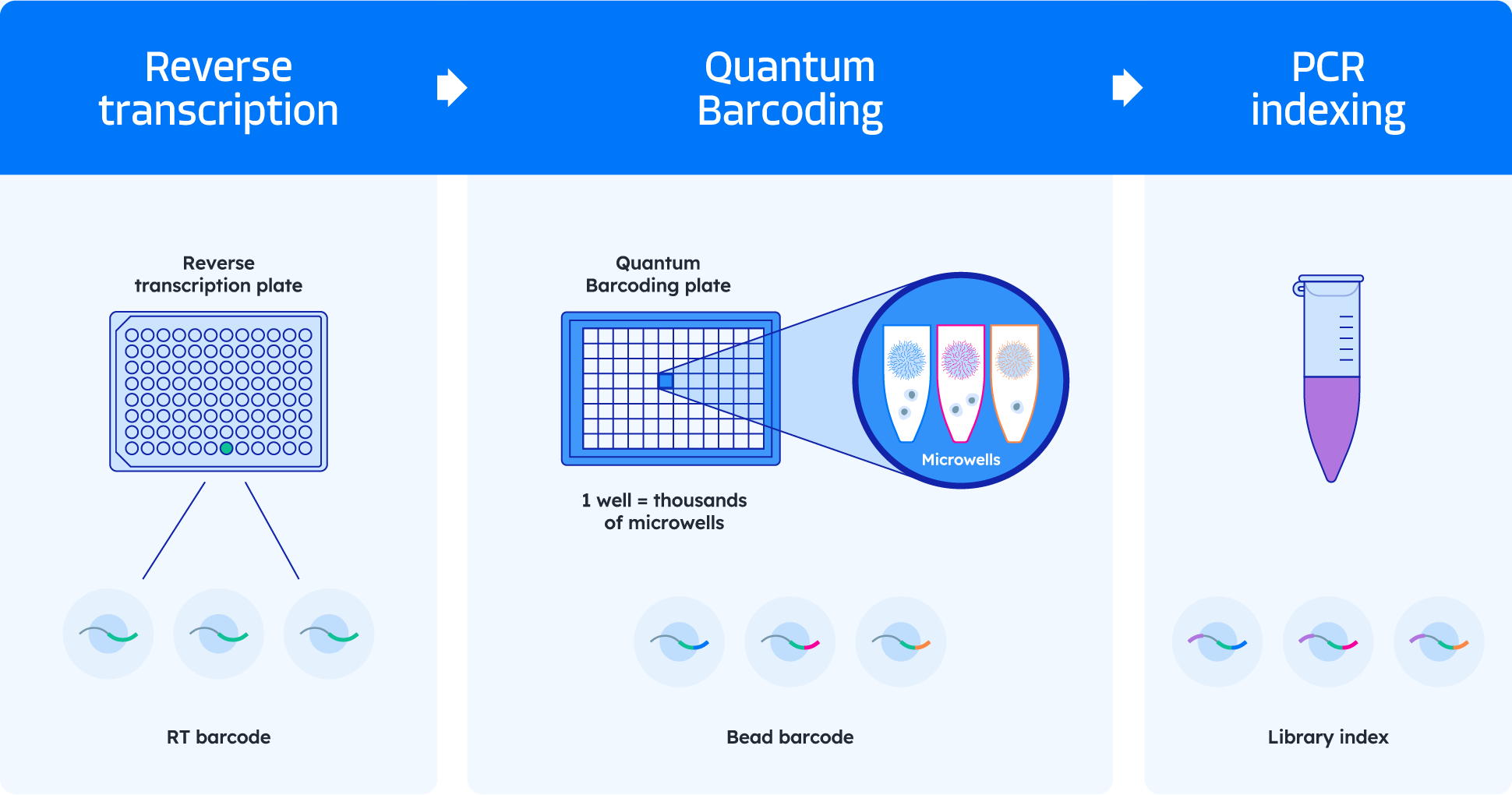
Quantum Barcoding technology dramatically streamlines your workflow
Quantum Barcode technology consolidates many levels of barcoding into one streamlined operation, cutting hands-on time by over 75%.
High performance for every scale
- Maximize insights with high transcript sensitivity
- Achieve cell recovery of 50%-60% or higher while keeping multiplets low (≤4%)
- Run projects of varying sizes with confidence using the same underlying technology
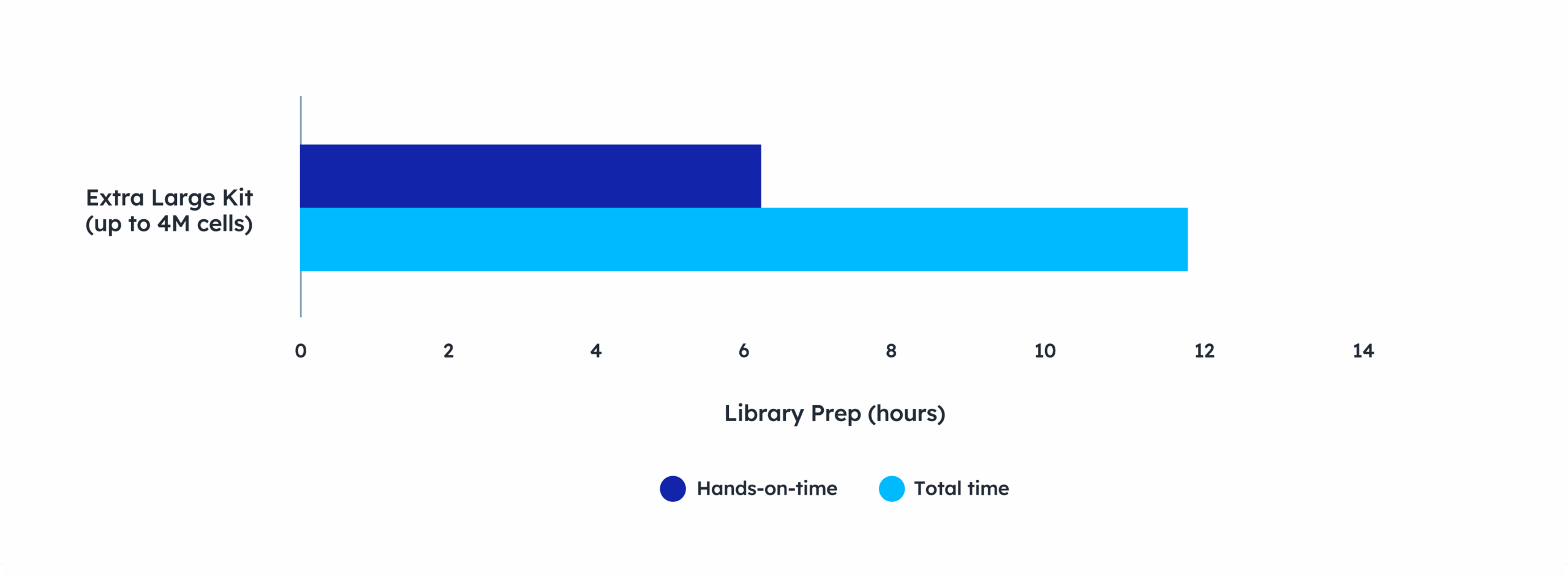
Lab efficiency for every scale
- Go from sample to sequencer-ready library for any sized project in 1.5 days
- Whether you are processing a few or thousands of samples, only half a day of hands-on time
- Easily scale projects with minimal increase in workflow time
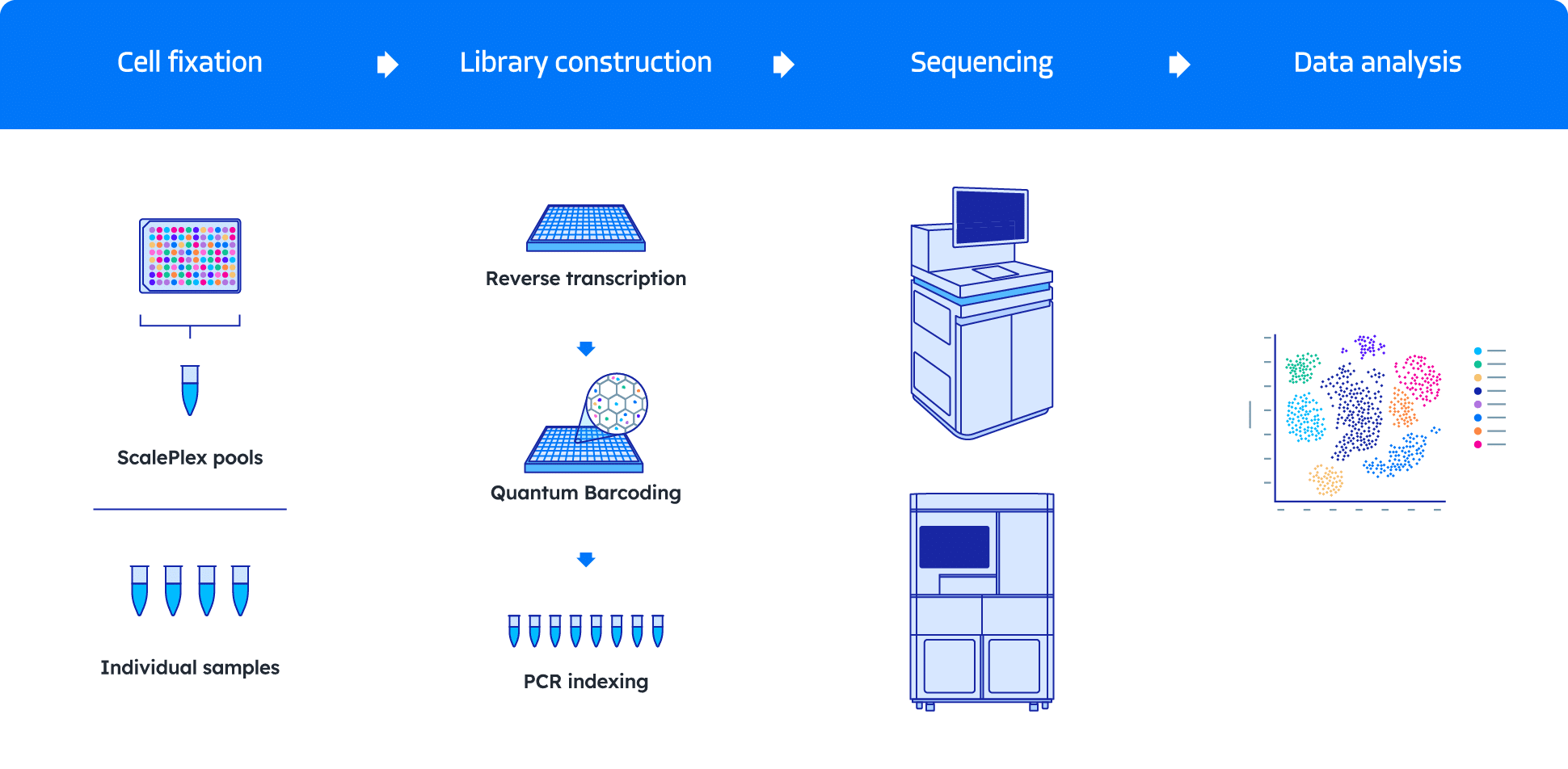
One streamlined workflow for every scale
Our comprehensive QuantumScale Single Cell RNA Sequencing workflow includes optimized sample fixation, library preparation using Quantum Barcoding technology, and integrated data analysis to power your research insights — all without requiring specialized equipment.
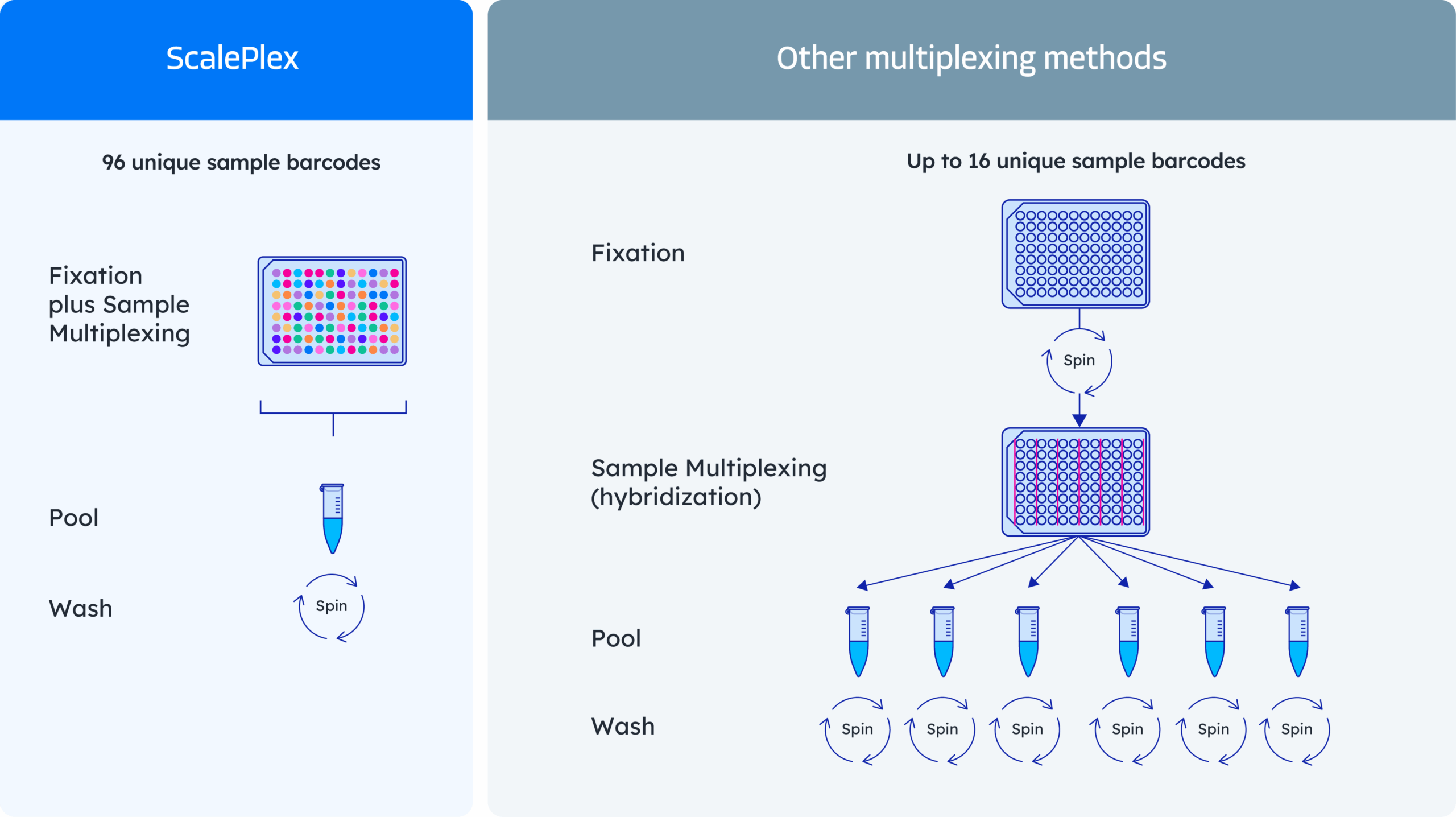
Effortless sample multiplexing for every scale
- Profile thousands of samples per run using ScalePlex
- No upfront optimization and compatible across species and cells or nuclei
- Simultaneously fix and label up to 96 samples in parallel, saving hours of time
- Minimize sample loss with fewer processing steps
Cost efficiency for every scale
- Flexible kit options offer the most cost-effective solution per cell, sample, and project
- Our technology enables exceptional value across a broad range of projects – from clinical research studies with a few samples to high-throughput screening with thousands of samples
- Dramatically reduce costs to under $100/sample when running hundreds of samples, making large-scale experiments achievable
- Profile thousands of samples per run using ScalePlex
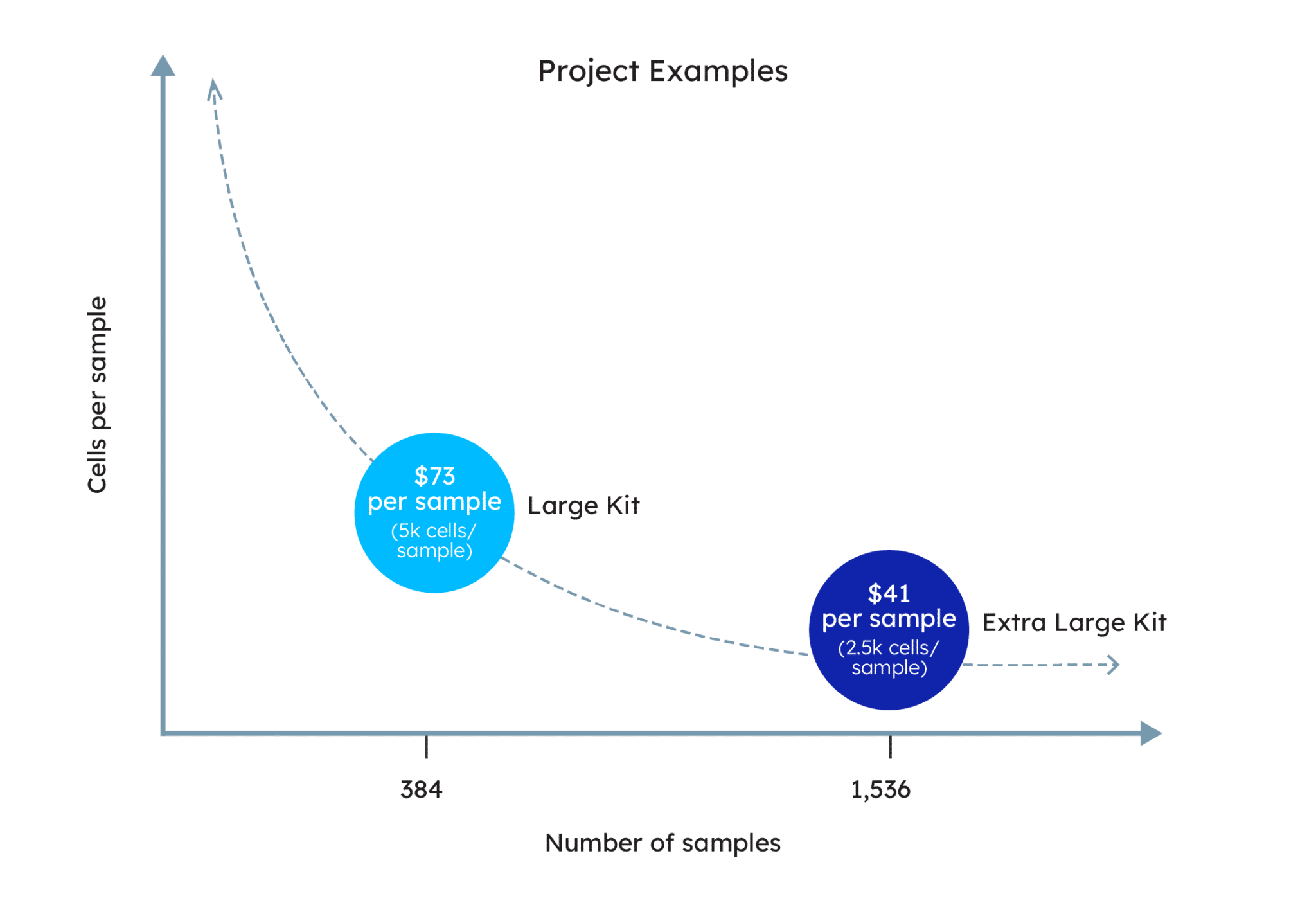
*Cost per sample includes fixation, sample multiplexing, and library preparation reagents
Kit size:
Choose the QuantumScale kit for your project:
Method
3' Whole Transcriptome
Number of samples at 10k cells/sample
16
200
400
Project run size (cells)
168K
2M
4M
Consider this kit for
Standard-sized projects
Biobanked specimens
Time-course or multi-site studies
Biobanked specimens
Time-course or multi-site studies
Biobanked specimens
Sample types
Cells, nuclei, frozen tissue, OCT-embedded samples
Any species
Sample number
1-24
1-96
1-96
Sample number with ScalePlex
Up to 2,304
Up to 9,216
Compatible flowcells
NovaSeq X - 1.5B or 10B
UG 100™
NovaSeq X - 25B
UG 100™
NovaSeq X - 25B
UG 100™
Frequently asked questions
We have demonstrated performance across a broad range of sample types, including but not limited to: tissue-dissociated cells, nuclei isolated from frozen and fresh tissues, PBMCs, and cell lines.
After fixation, samples can be stored immediately at -80C or transported on dry ice and then stored at -80C. Current tested storage recommendation is up to 1 year.
Samples can be fixed individually or fixed and pooled with ScalePlex oligos for streamlined multiplexing. Samples or ScalePlex pools can be processed immediately or stored for future use. When ready to start the library preparation, each sample or sample pool can be distributed across multiple RT wells for deeper profiling, or one RT well for maximum sample throughput.
Scale Bio provides a data analysis pipeline, the Scale Bio SeqSuite, which is compatible with sequencing instrument output files. The pipeline will de-multiplex samples, generate a cell-gene matrix, as well as provide a data quality metrics summary to assess run performance. The output files are compatible with many community software tools, including Seurat.
QuantumScale Single Cell RNA Sequencing libraries have been demonstrated with Illumina’s NextSeq and NovaSeq instruments, and Ultima Genomics’ UG 100™.
Yes, you can access our public data sets at the following link: ScaleBio Public Data Sets

Want to learn more about our QuantumScale Single Cell RNA Kit?
Reach out to our Team for more information or to request a quote.
Related products